Ask Almost A Doctor: Peptides, The Future Of Surgeons, And Viruses Causing Chronic Disease
Edition One
This is the first edition of what I hope will become a staple for Core Memory. Though I’d love to comment on politics, religion, culture and dating, I think my experience is best directed toward addressing questions about biology, medicine and healthtech. The vision here is to get you up to speed quickly on disparate bio and health topics through our regular installments.
For readers new to my stuff, I’m a 4th year medical student at the University of Vermont with prior experience in biological engineering of mosquitoes at George Church’s lab at Harvard, AAV (Adeno-associated virus) at Dyno Therapeutics and wearable monitoring devices at Caltech.
If you have questions, you can email me at eryneym@gmail.com, DM me on Twitter or Substack. Or put them in the comments below!
Also, none of the below constitutes medical advice. (Seriously. This is not medical advice - Ed.)
Enjoy.
Effie Klimi @effiebio
What do you advise people who have leaned onto the peptide craze? What will medicine look like if this trend keeps increasing in intensity?
Peptides, peptides, peptides . . .
I have written about these a bit with regards to the inflammation/pain claims that exist around BPC-157. My full thoughts on peptides are complicated, but can be summarized this way: buyer beware.
I think it is valid to point out that on-demand intelligence with AI or even just the internet has meant people feel like they can start taking control over their own health. You can really feel it, especially in tech communities as Jasmine Sun highlighted in NYT late last year. The problem is that humans are credulous, and hucksters know it. There is some balance to be had between four phases of trials, which include thousands of patients over 15 years, and me tweeting that Chemical X discovered by an Uzbek scientist in 1844 cured my face blindness. Unfortunately, right now things look too much like the latter than the former.
There is a very real, competent minority of well-resourced people who are looking to take healthcare into their own hands. Peptides are an example of that, and self-designing mRNA vaccines is another. In both instances, it is possibly the case that many N-of-1 treatments exist that could work for the individual, but there isn’t really any rigor that exists that can help generalize these results. That’s really the problem. Trial abundance isn’t my wheelhouse, but Rux Teslo has written extensively about clinical trial abundance, and I recommend you check that out.
Just so no one can say I am being completely unreasonable, I think that some of the peptides that people are buying actually do work. Specifically, I’m referring to retatrutide, Eli Lilly’s new triple hormone agonist that has shown better safety and efficacy than current GLP1s for weight loss. People are buying that one because they have (probably correctly) ascertained that the FDA’s stamp of approval is as good as guaranteed in a year or so. I just hope they reward Eli Lilly for their labor and actually get on that one when it comes out, instead of buying from a random compounder that is stealing EL’s IP.
If you’re into peptides, power to you, but consider asking your peptide dealer if they’ve run even a small blinded trial of 25 people. Or better yet, be agentic and organize one yourself. I firmly believe there is simply too great a cost to abandoning rigor.
Alex Kesin @alexkesin
Is the whole toxoplasmosis <> enhanced risk taking behavior thing real? Can I become a better poker player if I infect myself with it?
A myth has been created around the Toxoplasma gondii parasite. As far as I can tell, it stems mostly from work that comes from one Czech scientist Jaroslav Flegr. Throughout the 90s and 2000s, he started publishing increasingly provocative articles linking toxoplasmosis with high risk behaviors through surveys and questionnaires. According to him, those infected with the parasite are more likely to get into traffic accidents, be schizophrenic, and more recently, be entrepreneurs.
The problem is that there is a fairly iron-clad rebuttal from a group at Duke. They followed 1,000 people in New Zealand from birth until age 38, testing them for antibodies against the parasite to see who had been exposed or was currently infected, then tried to search for associations. Unfortunately for you, Alex, none were found.
Flegr himself is listed as a co-author on a study challenging the ground truth behind some of his earlier toxo<>high risk claims. In that paper, they tested for an association between toxoplasma infection and financial decision-making using real monetary incentives in a case-control design. They found no significant evidence that risk attitude or loss aversion was associated with infection.
There are lots of examples of mainstream media picking up tidbits of the most interesting, spurious findings in fields. You can find volumes of pop culture articles about how wine makes you live longer or how chocolate is good for you. Being a good citizen scientist means developing some antibodies against these sorts of studies. You’ll live longer that way.
Adith Arun @aditharun_
What does the future of being a doctor look like in the age of AI? Non-surgical only.
I wrote about five different answers to this question, so consider that my base level of confidence in the future. My current view has been shaped by discussions with Joe Janizek, a radiology resident at Stanford with a PhD in computer science.
The kinds of AI that will slowly encroach on core aspects of medical care are reliant on data for the most common presentations, and are therefore very good at managing them. Note-writing, diagnosis of simple rashes and prescription refills fall mostly under that. What happens with the most challenging stuff, though? I’m referring to those 1-2 patients you see who are just kind of confusing, where the story and medical picture just don’t make sense, and when you have a diagnosis, management is a mild nightmare. After talking to Joe, I think it is reasonable to assume the bulk of medicine will revolve around dealing with those cases exclusively. It’s not too dissimilar to what LLM-powered coding agents are doing for AI research right now, honestly.
If you are OK with having to keep your brain CPU at 100% all the time, then welcome to the future of medicine.
David Dales @d2dev_
What is the most exciting result you’ve seen happen to a patient lately (avoiding theoretical science)?
Antisense oligonucleotides are showing promise. One very recent win is Zorevunersen, a treatment for the devastating neurological disease Dravet syndrome, described in this New England Journal of Medicine article earlier this month. There is much I can say about the science, but I will instead just point you to this video. This is the type of result that makes me glad I went into biology and medicine. It’s undifferentiable from magic.
John Whittaker @johnowhitaker
Microfluidics in bio: buzzword or are there lots of genuine good uses? My mind buckets ‘microfluidics’ with ‘graphene’ as something hyped as having lots of potential solutions that are always somehow a little ways in the future...
I don’t know much about graphene, but I think it’s clear that microfluidics has already made a real impact on biology: single cell RNA sequencing. It’s advanced enough since 2013ish that 10X Genomics licensed some of the original technology out of Harvard and now anyone in the US can implement it in their workflow. With 2025 revenue of nearly $700m, we can count this as a win for microfluidics. If digital droplet PCR is included in this category of boring quantification tools, the revenue goes well past $1B annually. For reference, the graphene global market across all companies is less than the revenue of a single company using microfluidics for a single assay.
There are other areas where I think the graphene comparator is a little more true, though. Organ-on-a-chip depends on microfluidics, but the technology has yet to really break out of labs. The FDA (and the Wyss Institute) are hoping that this ends up being the thing that can move the biotech industry away from reliance on animal models, and are legislating as such. Time will tell.
In my mind something stops being a meme once it becomes so boring and integrated into a field that it no longer becomes associated with a buzzword. I don’t know many people losing their minds over droplet quantification.
Adic @adic_9
Do you think we’ll get broadly available CAR-T or similar autoimmune drug free remission meds in the next 10 years?
CAR-T is a new cell therapy modality that mostly gets paired with cancer, but there’s reason to be fairly optimistic it has potential for autoimmune disease. I’ll point specifically at lupus. A case series a couple of years back involving a cohort of severe lupus and some other patients showed a CD19-targeted CAR-T treatment resulted in 100% disease-free remission. This persisted for the duration of follow up, averaging 15 months across the patient cohort.
Open questions remain about how durable these treatments are, but, on the surface, there’s reason to be excited. As with all things frontier, cost is a pretty big burden, but the current mainstay of autoimmune diseases – monoclonal antibodies – ain’t exactly cheap either, and patients are getting those at least monthly. I’m optimistic on this one. I put my p(win) at 75% by 2035 for an SLE treatment clearing a Phase III clinical trial.
Niko McCarty @NikoMcCarty
Why is the experimental hit rate for protein binder design so low?
Deep ball knowledge from Niko here. Proteins are dynamic. Everyone knows this, but what can really be done with this information? Not much right now.
AlphaFold and the tools that followed it made it easy to model static proteins, but don’t perform so well with proteins that move. There’s a neat paper from 2018 that suggests that as much as 5% of all proteins known in the Protein Data Bank are fold-switching, meaning they have extreme conformation changes in their active vs inactive state. If you used AlphaFold, you might get one state or the other, but, with no way to see the full spectrum of conformations, it’s hard to know how some proteins engage with their target.
This is doubly true for things like antibody binders, where the interaction requires understanding how a designed binder changes between its states. Thinking about antibodies, complementarity-determining regions (CDRs), particularly the H3 loop, are among the most conformationally diverse structural elements in all of biology. They’re essentially floppy loops that sample a huge ensemble of states, and the binding-competent conformation may only be transiently populated. So you’re trying to dock two moving objects against each other, and the computational tools are handing you a single snapshot of each.
And this gets at a deeper problem with how the field currently designs binders. Most pipelines optimize for a single, static interface. They score a candidate based on how well it packs against one conformation of the target, with maybe some light sampling around the backbone. But the protein your binder actually encounters in solution, on a cell surface, or in an assay isn’t frozen. A designed interface that looks perfect against the crystal structure may be competing with a conformation that buries the epitope entirely, or that rearranges key side chains at the binding site. You don’t see that with RosettaFold or Chai or Boltz.
I don’t want to point any fingers, but I’m afraid part of why there seems to be a disconnect between the papers and the reality stems from the way binder benchmarks are highlighted in papers. State-of-the-art generative design tools typically report experimental hit rates in the single-digit percentages for novel targets because they specifically select from carefully chosen, well-behaved targets with rigid binding sites. It doesn’t work as well when you move to a GPCR, a cytokine receptor with a flexible extracellular domain, or a viral glycoprotein that samples multiple perfusion states. It’s basically guaranteed that your hit rate drops, it’s more a question of whether that’ll be 10-fold or 100-fold.
A second, more nuanced perspective is that the energy landscape of binding is shallow. The difference between a binder with nanomolar affinity and one that doesn’t bind at all can come down to one or two residue contacts worth of free energy (~1–3 kcal/mol). Current models, even good ones, don’t resolve energy differences at that scale reliably, meaning they have very poor resolution for single mutation variants. When your scoring function has noise on the order of the signal you’re trying to detect, you’re essentially gambling on which designs to take to the lab.
The solutions to these problems are pretty boring. Mostly it comes down to finding a way to unify dynamics with the current frontier models. Molecular dynamics tries to get at this, but they’re a PITA to use and have their own limitations. I’m sure someone’s working on this. Once that’s solved, I expect paper hit rates to become real.
Niko McCarty @NikoMcCarty
What is the connection between certain viruses (ie Coxsackie) and various chronic diseases? Is this understudied?
The broader virus-chronic disease link is quite well-established at this point, so it feels odd to say it’s understudied but, well, it’s understudied. A current well-described example of one such virus is Ebstein-Barr Virus (EBV) better known to the public as the cause of mono. EBV is involved in a ton of processes. It can cause nasopharyngeal carcinoma, certain lymphomas, and more recently in 2022, was shown to be a major driver of multiple sclerosis. The MS component is important because of the amount of effort required to show the dynamics.
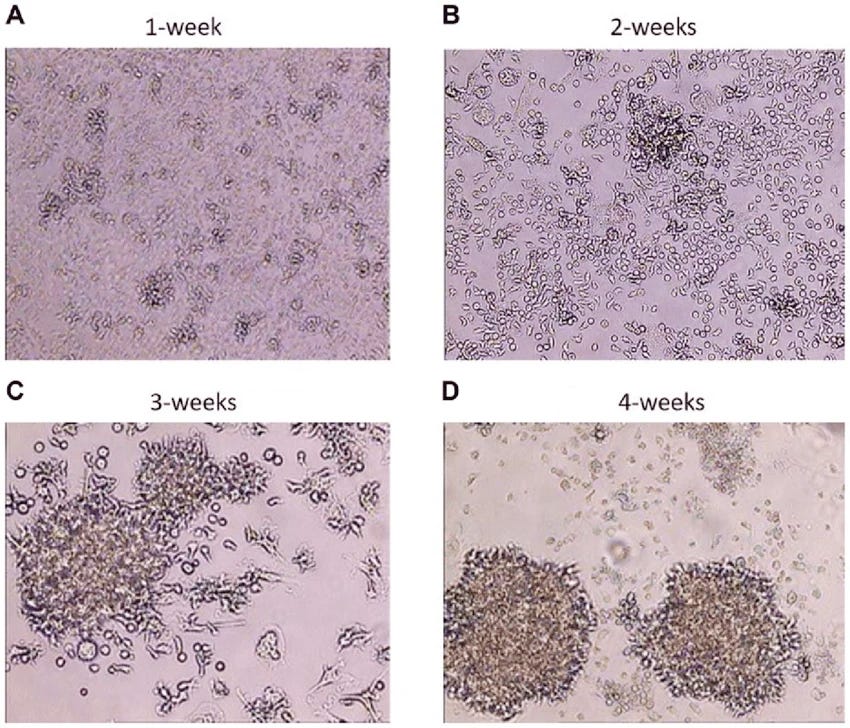
A Harvard group used millions of patient samples to demonstrate that EBV is essentially a prerequisite to developing MS, whereas the associations I mentioned before took significantly less effort. A component of that seems to be related to the fact that the kind of data required to demonstrate causality for disease processes like MS are molecular in a way that was not needed for cancer. For instance, for lymphoma, you can actually see, visually, the cells infected with EBV change their appearance and behavior. If you look for long enough, you’ll see them turn. Maybe MS has a similar thing, but we don’t know it yet and there’s no way to sample from the brain easily, so large data cohorts are required for now.
Time course of EBV infection. Doesn’t require being a doctor to see that something is going on there.
There are plenty of other examples here, by the way. T-cell leukemia/lymphoma is caused by HTLV-1, another virus, while hepatitis C can cause liver cancer if someone is infected chronically for long enough. Just last year there was the result that the shingles vaccine seems to stave off dementia (though maybe some selection bias was involved). I don’t have any strong opinion on the chances this will happen with Coxsackie virus, but it would not surprise me if it turns out to be implicated in some autoimmune processes, especially since there’s some evidence that it is related to type 1 diabetes. It seems like the challenge is mostly about finding the thing to anchor on when looking at large cohort studies, as was the case with EBV x MS.
Jonas Kubilius @qbilius
Top-3 diseases that could be tackled if we came up with novel (epi)genome editors (not just more prime editors)
You said three diseases, so the easiest answer is all diseases of the same mechanism: Huntington’s, myotonic dystrophy, and Fragile-X syndrome. They’re all caused by trinucleotide repeats that are hard to address with existing prime editors. What you would ideally want is a way to silence expanded alleles while preserving safe copies – a task that lends itself well to epigenetics.
Imprinting disorders like Prader-Willi or Angelman seem viable here, too. They are caused by unintended silencing of either the maternal (Prader-Willi) or paternal (Angelman) copies of chromosomes. Reversing that cannot be done with gene editors, but can theoretically be done with epigenome ones. I’m less optimistic that you can undo the developmental consequences of that imprinting, though.
Jonas Kubilius @qbilius
In one of your pieces/podcasts you were skeptical of personalized genome editor costs ever going down below 100k. Why is that?
To clarify my position, I view this as more of a delivery problem than an editor one. The core issue is that even if you have a perfect genetic cargo that can solve a particular disease, we don’t have great tools that reliably get it all where you need it to. Delivery is just not a solved problem, with the exception of the liver where cheaper options like lipid nanoparticles are viable. For every other organ, the best tools we have are viruses, and until gene therapies can move beyond either natural or synthetic viruses, I just really don’t see how the cost can go below $100,000. The reagents alone, ignoring regulatory costs, for an adult human sized batch of AAV or lentivirus might cost you $50,000 minimum. It’s likely way more if avoiding immediately killing someone with endotoxin from your production process is part of your objectives.
It’s not free to synthesize the input DNA required for human-sized batches of virus, and your reagents have to be actually sterile, which comes at a pricing premium. This is a place where cost saving on reagents makes you penny-wise but pound foolish. Throw in costs associated with trials and you’re talking about several hundred thousands more to the cost of the treatment, and that’s just to break even.
I do sincerely hope other, cheaper delivery modalities can pop up that give tissue-level resolution, but that hasn’t happened so far. For now, viruses are what we have. Everything comes at a cost.
Mosasaurus @mosasaurus27
Is it accurate to view/represent the cell as a machine?
No, it is not accurate, though it is convenient to say so. Unlike machines, cells are never off unless they are dead. Processes are dynamic. Pathways do not pause, but rather just slow down. Everything is happening chaotically all the time, with inputs and outputs feeding into their own pathway and kicking off other processes, too. A neural network feels like a slightly better representation of how a cell behaves, which means that computation can serve as a tool for modeling cells. But an actual machine, no.
My old colleague Duo Peng and his team at CZI have some interesting work towards establishing a better model for how a cell behaves, but it’s not quite there yet.
Ashlee Vance @ashleevance
What’s the deal with the dog cancer thing?
Rosie and The Australian Cure is an interesting case of all discourse simultaneously being right and wrong, my own contributions included. If I had to summarize, the current stances are scientists saying the treatment doesn’t work, and the tech world saying they’re missing the point, and this is all about one man’s extreme agency being turbocharged by AI. On the surface, it’s the story of an Australian entrepreneur using his financial resources and AI to act outside of the healthcare system to treat his beloved pet of her cancer. Heartwarming. Sadly, misleading.
According the reporting we have so far, Rosie’s owner used ChatGPT to design an mRNA vaccine against her solid tissue tumor, resulting in a 50% reduction in size. The use of consumer AI to troubleshoot illness is cool in its own right, but the trouble comes when techno-optimism pushes the suggestion that the cure to cancer is at our fingertips thanks to Grok. Truthfully, there’s reason to want to pump the brakes a little bit here. Namely, we don’t even know if the guy’s treatment even did anything because he simultaneously treated his dog with a checkpoint inhibitor, which has already been established as a treatment for her cancer type. The technical details are scant as of now, but the ones we do have, especially around the use of AlphaFold in the process, also has me scratching my head a bit.
I completely understand the desire for optimism. It is empowering to believe that on-demand intelligence will allow everyday people to address the problems in their lives, but it’s just not the case that AI added much here. If you’re looking for examples of that being true, there are better stories that champion the use of AI in giving laymen the capabilities of the scientific elite – even examples in humans! Check out the story of Sid, who used his resources and AI to treat his own cancer.
Why the need to stretch the truth? I suspect a part of it is a desire to believe that we can completely discard the shackles of establishment science. Peptides have shown that people feel they are as capable as doctors and scientists when it comes to their own health. I think this is just another instance of that. But unlike the inflammation meme perpetuated by Elite Human Capital of SF, which is embarrassing but mostly harmless, creating false hope for diseases like cancer is horrible. It’s actually worse than horrible, it’s shameful, and I think caution is in order.
People with limited time left get desperate and justifiably reach out to grab everything and anything they think will help them. Abandoning mechanisms to separate the wheat from the chaff – which is what clinical trials are meant to do – will mean condemning people who picked wrong.
Let’s see how this story plays out and revisit it in a couple months. As I see it, there are enough scientific gaps that call for skepticism.
Happy Friday.